



Tierras, Geomorfología y Recursos Naturales
INFLUENCE OF EDAPHIC/CLIMATIC FACTOR ON MICROBIAL POPULATION IN ARID SOILS WITH IRRIGATED AGRICULTURE
INFLUENCIA DE FACTORES EDAFO/CLIMÁTICOS SOBRE POBLACIONES MICROBIANAS DE SUELOS ÁRIDOS CON AGRICULTURA BAJO RIEGO
Boletín Geográfico
Universidad Nacional del Comahue, Argentina
ISSN: 0326-1735
ISSN-e: 2313-903X
Periodicity: Semestral
vol. 43, no. 2, 2021
Received: 25 November 2020
Accepted: 28 December 2021
Abstract: The agricultural frontier over the arid marginal zones is steadily expanding by means of irrigation technology. Considering the impact of bacterial population and its biological activity on soil homeostasis, information on soil biodiversity and its biological activity are a significant input to maximize the production with minimal impact on these susceptible environments. The aim of this exploratory work was to assess the effect of long term agricultural practices (30 years) on the activity and structural aspects of the microbial community from arid irrigated agricultural soils. For this, three environments were evaluated, collecting samples of a Pristine arid soil (without irrigation and undisturbed) and two different irrigated agricultural (Alfalfa and Corn) throughout the 2014-2015 season. For this, samples from Pristine (without irrigation) arid soil and (Alfalfa and Corn) were collected along the 2014-2015 season. Soil physicochemical analysis was performed, temperature and humidity were recorded, as well as total biological activity (FDA hydrolysis test) and molecular fingerprinting (DGGE and T-RFLP) for each treatment. Soil physicochemical analysis show that agricultural practice improved the edaphic properties like organic matter level, although the contribution of water in limited quantities (pressurized irrigation) an increase in electrical conductivity. Total biological activity along the seasons showed similar profiles for all scenarios, and the coldest/unirrigated and hottest/irrigated-rainy months showed the lowest and highest activities respectively. Overall, the Pristine condition presented significantly lower biological activity levels along the season. The DGGE fingerprint showed a discrete pattern with low number of bands and, against expected, showed no differences between the three scenarios. Subsequent T-RFLP analysis confirmed that soil bacterial biodiversity was not modified by decades of intensive irrigated agricultural activity.
Keywords: Biological activity, Irrigation, Arid ecosystems.
Resumen: Las fronteras agrícolas se están expandiendo constantemente, por medio de la tecnología de riego, hacia zonas áridas marginales. Considerando el impacto de la población bacteriana y su actividad biológica en la homeostasis del suelo, la generación de información sobre la biodiversidad del suelo y su actividad biológica son un importante aporte para maximizar la producción con un impacto mínimo en estos ambientes susceptibles. El objetivo de este trabajo exploratorio fue evaluar el efecto de las prácticas agrícolas a largo plazo (30 años) sobre la actividad y los aspectos estructurales de la comunidad microbiana de los suelos agrícolas áridos con agricultura bajo riego. Para ello, se evaluaron tres ambientes, recolectando muestras de un suelo árido Prístino (sin riego y sin disturbar) y dos suelos agrícolas bajo riego con diferente historial de manejo (Alfalfa y Maíz) a lo largo de la temporada 2014-2015. Se realizó un análisis fisicoquímico de los suelos, se registró la temperatura y humedad, así como la actividad biológica total (prueba de hidrólisis FDA) y la huella molecular (DGGE y T-RFLP) para cada tratamiento. Los análisis fisicoquímicos del suelo muestran que las prácticas agrícolas mejoraron algunas de las propiedades edáficas como el nivel de materia orgánica, aunque el aporte de agua en cantidades limitadas (riego presurizado) provocó un aumento de la conductividad eléctrica. La actividad biológica total a lo largo de las estaciones del año mostró perfiles similares para todos los ambientes, y los meses más fríos/sin riego y más caluroso/con riego mostraron las actividades más bajas y más altas, respectivamente. En general, la condición Prístina presentó niveles de actividad biológica significativamente más bajos a lo largo de la temporada. La huella dactilar DGGE mostró un patrón discreto con bajo número de bandas y, contra lo esperado, no mostró diferencias entre los tres ambientes evaluados. El análisis por T-RFLP posterior confirmó que la biodiversidad bacteriana del suelo no fue modificada por décadas de intensa actividad agrícola irrigada.
Palabras clave: Actividad biológica, Riego, Ecosistemas áridos.
Introduction
The irrigated agriculture is a key strategy to produce crops in arid and semiarid regions. In this way, extension of land under irrigated agriculture has steadily grown in developing countries from 47.3 million hectares in 1900 to 276.3 million in 2000 (Matson, Parton, Power & Swift, 1997; Scanlon, Jolly, Sophocleous & Zhang, 2007). Therefore, considering the yields obtained from irrigated agriculture, in comparison to the unirrigated one, it is expected that irrigated areas will continue increasing by 20% until 2030 (Tilman, 1999; Scanlon et al., 2007).
In arid and semi-arid soils, the water deficit is permanent. This condition is generated by the combination of low precipitations and the elevated evapotranspiration caused by strong solar radiation. In this context, the use of irrigation systems combined with heat stress tolerant crops are the alternatives for sustainable agricultural production (Evans & Sadler, 2009).
At the same time, there is evidence of the hydrological impacts generated by the conversion of natural systems into irrigated ones, such as the elevation of the water table (Pérez, Abrahao, Causape, Cirpka & Burger, 2011), the increase of evapotranspiration and the redistribution of salts and nutrients in the soil profile (Scanlon et al., 2007; Gabriel, Almendros, Hontoria & Quemada, 2012). Therefore, even providing enough water for a maximum crop development, it must overcome other disadvantages, such as low fertility and salinity of soils.
Microbial communities play a vital role in maintaining soil health in the especially fragile arid and semi-arid ecosystems. In this way, soil microbiota mediates 80–90% of soil processes, including nutrient cycling, decomposition of organic matter, carbon sequestration, soil genesis, soil structuring and consequently water holding capacity (Singh et al., 2006; Zhao et al., 2014). These processes help to sustain soil resilience for recovering from perturbations (i.e. topsoil systematization), essential for a sustainable crop production in arid and semi-arid environments (Griffiths, Bonkowski, Roy & Ritz, 2001).
It has been reported that soil characteristics and plant diversity strongly influence the function and structure of the microbial community (You et al., 2014). Thereby, global change or anthropogenic intervention such as agricultural practices, affect the composition, abundance and biological activity of the soil community (Matson et al. 1997; Francioli, Ascher, Ceccherini & Pietramellara, 2014).
On the other hand, considering microbes have shorter turnover rates than plants and earlier reflect the changes in land management, they can be used as sensitive indicators of anthropogenic activities (Entry et al., 2008). Since it is a simple, sensitive and economic measurement tool, soil enzymatic activity results a reliable indirect indicator of changes in soil ecosystems (Meena et al., 2014).
In addition, changes in the microbial community structure can be directly assessed by culture-independent and quick methods for fingerprinting microbial communities like 16S rDNA Terminal Restriction Fragment Length Polymorphism (T-RFLP) and Denaturing Gradient Gel Electrophoresis (DGGE) (Enwall & Hallin, 2009; Francioli et al., 2014). These approaches have provided a more thorough insight into changes occurring in the structure of the microbial community as a whole (Enwall & Hallin, 2009). In this way, the relative abundance of common species present in a sample can be determined, independently of the conventional cultivation methods.
In this context, the aim of this exploratory study was to assess the effect of agricultural practices on the activity and structural aspects of the microbial community from irrigated agricultural arid soils. Thereby, microbial activity of soils with different management history and the structure and biodiversity of their microbial communities were compared by using FDA hydrolysis test and DGGE combined with-RFLP respectively.
Materials and Methods
Site description.
The sites selected are agricultural lands dedicated since the 1980 decade to crop production under irrigation, located near to 25 de Mayo, La Pampa, Argentina (37º 54´ 09.58”S 67º 45` 55.12”O). This region presents an arid continental climate, characterized by cool winters and hot summers with scarce annual rainfall (mean 250 mm). The rain season is from October to March, with an average annual temperature about 15.0 ºC.
Soils were classified as Entisols (Panigatti, 2010). The main feature of these soils is the little or no evidence of pedogenic horizons development and reduced level of organic matter (OM) as product of large alluvial deposition of sand and little level of clay (del Valle, 1998). Moreover, due to their origin, these soils have a basic pH and are extremely heterogeneous; all those characteristics defined them as arid soils, with low fertility and high permeability (Panigatti, 2010)
Experimental design, soil samples collection and characterization.
Different agroecosystems were considered to evaluate the effects of the local agricultural management on the soil microbial population. Selected sites included a corn field (Zea mays) under gravitational irrigation since 1987, an alfalfa field (Medicago sativa) under pressurized irrigation since 2002, and finally a characteristic open natural chaparral with a few vegetation; regarded as Corn, Alfalfa and Pristine soil respectively.
The samples were collected at 0–10 cm depth, at seven deferent times between March 2014 and January 2015 using a soil auger. Every sampling time, six composite samples were collected from the Corn plot, twelve composite samples from the Alfalfa plot and four composite samples from the natural undisturbed chaparral (pristine soil) according to soil heterogeneity of each plot. The sampling points were randomly chosen and georeferenced to sample the same spot along the experiment. Samples were transported to the laboratory in a plastic insulated container at 4.0ºC and immediately processed. Parallel, a part of each sample was reserved for DNA extraction at -20.0ºC until the moment of use.
During sample collection, soil moisture and temperature was recorded using an ECH2O Check soil moisture monitor (Decagon Devices) and a digital thermometer probe, respectively.
Soil physicochemical análisis
Bulk soil collected from the three different plots was processed to determinate the particle size by the Bouyoucos hydrometer method (Bouyoucos, 1962), the organic carbon content by the Walkley and Black procedure (Walkley & Black, 1934), the electrical conductivity (EC) from soil saturation extract and pH 1:2.5 (1 soil: 2.5 water relation). Both, EC and pH where measured whit the bench top pH-mV-Conductivity-TDS- Meter Altronix mod. EZDO-PC. Specifically, to pH measurement it was utilized a PY41 Alpha Electrode and to EC a platinum Altronix probe.
Soil enzymatic activity test (FDA hydrolysis).
The total microbial activity was determinate according to Adam & Duncan protocol (Adam & Duncan, 2001). Briefly, two grams of soil (fresh weight, 2 mm sieved) was placed in a 50 ml conical flask and 15 ml of 60 mM potassium phosphate buffer pH 7.6 were added. Then, 0.2 ml of 3´6´-diacetyl-fluorescein stock solution (1000 mg FDA ml-1) were added to start the reaction (3´6´-diacetyl-fluorescein, Sigma-Aldrich Co. Ltd). The flasks were stoppered, shaked and placed in a water bath at 30.0 °C with constant stirring (100 beats.min-1) for 20 minutes. Parallel, blanks were prepared without the addition of the FDA substrate along with a suitable number of sample replicates.
After incubation, 15 ml of chloroform/methanol (2:1 v/v) were added to terminate the reaction. Stoppers were replaced and the contents thoroughly shaked. The contents of the conical flasks were then transferred to 50 ml centrifuge tubes and centrifuged 3 min at 2000 rpm. The supernatant from each sample was then filtered (Whatman, N° 2) into 50 ml conical flasks and the filtrates measured at 490 nm (Metrolab 1600 Plus Ver 3.04d spectrophotometer).
The concentration of the fluorescein released during the assay was calculated using a 0 to 5 mg fluorescein ml-1 standards calibration curve (Fluorescein sodium salt Sigma Aldrich 46960)
Extraction of Community DNA from Soil.
Total DNA was extracted from two sample poits to perform DGGE and t-RFLP fingerprint techniques. The plot choice to DNA extraction (Table 1) was made selecting the similar in percentage of clay plus silt (c+s) to minimize the textural effect on the edaphic community. In this sense, many authors have shown that texture influences the composition of the microbial community affects litter decomposition affecting soil water availability, pore size distribution, nutrient availability, and surface area (Fang, Kremer, Motavalli, & Davis, 2005, 2005; Hamarashid, Othman & Hussain, 2010; Bach, Baer, Meyer & Six., 2010). The total DNA was extracted (0.25 g of each sample) by means of a ZR Soil Microbe DNA MiniPrep kit (Zymo Soil, U.S.A) according to manufacturer specifications. Extracted genomic DNA was PCR amplified according to the protocols from DGGE and t-RFLP techniques (2.6 and 2.7 sections). It is important to highlight that we could only obtain one DGGE sample of corn, probably because the extracted DNA was not able to this technique.
DGGE: PCR amplification of the 16S rDNA gene.
For this technique samples: Pristine a, Pristine b, Alfalfa a, Alfalfa b and Corn a were used (see soil physicochemical analysis in Table 1). The bacterial 16S rDNA gene was amplified by nested PCR to increase de sensibility of technique according Solaiman & Marshner (2007), using the 27F-1495R and finally the pair 341F-GC/534 R universal primers (Ziesemer, et al., 2015). The PCR was carried out in 50 µl (final volume) using a Mastercycler epgradient (Eppendorf, Germany). Each second-round reaction was carried out by using 1 µl of PCR product from first round, in 2 µl of 5X PCR buffer, 2 mM dNTPs, 0,5 mM of each primer, 0.1 µl of Taq DNA polymerase (Promega, Madison, WI, USA (5 U/µl), using the following PCR profile: an initial denaturing step of 5 min at 93.0ºC, 29 cycles of 30 sec at 94.0ºC, 30 sec at 53.0ºC, and 2 min at 72.0ºC, with a final extension of 10 min at 72.0ºC. Amplified DNA products were checked by electrophoresis on 1% agarose gel, stained with ethidium bromide and photographed under a UV light (302 nm). Finally, amplified DNA products were resolved by denaturing gradient gel electrophoresis (40-60%) employing the Bio Rad D Gene System (BioRad, Hercules, CA, USA). The electrophoresis was performed at 60.0ºC in 1X TAE buffer at 100V along 20 min, then at constant voltage (70 V) during16 hours. The staining was made with a solution of TAE 1X buffer (Tris- Acetate-EDTA)/SYBR Safe 1X. The gel image was captured with Gel Doc 2000 (Bio-Rad).
Cluster analysis and diversity índices.
Profile and cluster analysis, and dendrogram generation were carried out using the BIONUMERICS® program version 6.6. The similarity between the band patterns wascalculated using the Pearson coefficient and the clustering analysis was performed with the unweighted pair group method with arithmetic averages (UPGMA) to generate a dendrogram by using mathematic averages algorithm programs integral to the BIONUMERICS® software. The coefficient of cophenetic correlation was used to measure the consistency of clusters.
The number of DGGE bands present in each sample was used to measure the Richness index (R). The Shannon-Weaver index of general diversity (H´) and the Simpson index of dominance (D) were calculated considering the number of bands and the relative intensities of each band (Pi) in each lane. Relative signal intensities of detected bands, in each gel track, were determined by using the Quantity One software package (Bio-Rad) and calculated from the peak area of the densitometric curves.
T-RFLP: PCR amplification of the 16S rDNA gene.
The bacterial 16S rDNA gene from the sample Pristine a, Pristine b, Alfalfa a, Corn a and Corn b were amplified by PCR using 27F-1492R universal primers (Weisburg, Barns, Pelletier & Lane, 1991). The indicator 27F is marked with FAM (6-carboxyfluoresceina) to visualize the terminal restriction fragments (TRF).The PCR reaction was carried out in 50 µl (final volume) with 10 μl 5X of buffer Go Taq (Promega), 20 mM on each dNTPs, 36.25 μl of ultrapure sterile water, 1 μl of each indicator, 0.25 μl of Go Taq DNA polimerase (Promega) and 1 μl of DNA template. The PCR profile utilized consisted in an initial denaturing step of 5 min at 95.0 ºC, 35 cycles of 30 sec at 95.0 ºC, 30 sec at 55.0 ºC, 2 min at 72.0 ºC and a final 5 min step at 72.0 ºC. Amplified products were checked by electrophoresis in agarose gel before digestion step. The TRF digestion was realized according to Promega with Hae III and Cfo I restriction enzymes. TRFs analysis was performed by the Department of Genomic (Biotechnology Institute, INTA Castelar) using capillary sequencers (ABI131 30xl Applied Biosystems). TRFs length and peaks were determined using the Peak Scanner software (ver 1.0). The consensus profile was generated by each sample using the T-Align software (http://inismor.ucd.ie/~talign/). This profile contains a list of TRFs and relative fluorescence intensities (Smith et al., 2005).
Data analysis and statistics.
Soil physicochemical, temperature and humidity parameters and total biological activity were analyzed by one-way ANOVA for a completely randomized design and a Duncan-Test post hoc (Duncan, 1955). DGGE band pattern was obtained in a GelDoc 2000 imaging system (BioRad) and results were analyzed by Dice similarity coefficient and Cluster method: UPGMA– Cophenetic index. The electropherograms were analyzed with PeakScan software and the fragment frequency interpreted by T-align (web-based tool for comparison of multiple terminal restriction fragment length polymorphism profiles).
Soil physicochemical properties
When the physicochemical parameters were analyzed (Table 1), the EC and total organic matter % (TOM%) presented the most relevant differences. Regarding the EC, the Alfalfa environment presented significatively higher EC than the other environments (p<0.05). This is most probably related to the salts contribution from irrigation water to the managed lands. In this way, although the efficiency in the water use from a central pivot (Alfalfa field) is significantly greater than the gravitational system (Corn field), Evans & Sadler (2009) pointed that the salts could be accumulated in the agricultural profile. Conversely, the irrigation sheet applied in the gravitational model in Corn plots is considerably greater than that supplied by pressurized irrigation. This excess of water could help, through leaching, to “wash” the salts and therefore to reduce the EC.
On the other hand, when the pH was analyzed, did not significant statistical differences were found. The Pristine conditions shown the highest pH values and the Alfalfa fiends the lowers, being the corn points those that resulted with an intermediate pH value. When relating the pH with the EC, it was observed an inverse relationship. In this sense, at higher pH, the EC was lower and vice versa, probably because soil pH may affect the solubility of salts. Soil EC relates directly to the presence of soluble salt in the soil. Alkaline soils, typical of arid regions, could have less amount of soluble salt and consequently lower EC values (Mohd-Aizat, Mohamad-Roslan, Sulaiman & Karam, 2014), being the agricultural activity the cause of the observed decrease in pH.
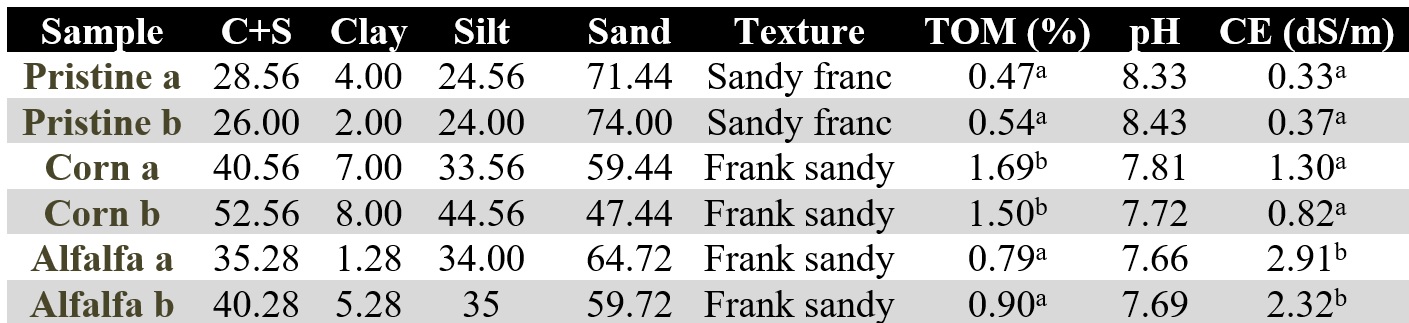
In general, the %TOM was low, although it was observed a positive correlation between the increment in %TOM and the long-term of the agricultural activity of each plot. In this sense, the unmanaged Pristine soil showed lowest level (0.6 % approximately) and Corn soil the significant higher value (1.5-2%) (p<0.05), in accordance with its 30 years of agricultural management. The alfalfa plots presented, after 15 year of agricultural management, intermediate values to the other two plots (0.8-0.9% approximately) (Table 1). The observed differences prevailed on spite of the eventual systematizations applied to the managed plots, especially on the gravitational irrigated Corn fields.
Total soil enzymatic activity.
The fluorescein diacetate (FDA) hydrolysis is an easy and fast method to assess the Total Biological Activity (TBA), used on a wide range of samples (Adam & Duncan, 2001; Green, Stott & Diack, 2006; Şumalan, Alexa, Negrea & Doncean, 2010). The enzymes responsible for FDA hydrolysis (Non-specific esterases, proteases and lipases) are abundant in the soil environment and have been shown to hydrolyze FDA; so that, could be involved in the decomposition of many types of tissue. Therefore, the ability to hydrolyze FDA seems to be widespread, especially among the major decomposers, bacteria and fungi (Schnürer & Rosswall, 1982).
By means of FDA test, the TBA from the three different environments (Pristine, Corn and Alfalfa) was assessed between March 2014 and January 2015 (Figure 1A). Although the TBA level was different at each site it was found that, on average, the TBA decreased during cool months, recovering in the months with higher temperatures. The seasonal variation observed in the TBA can be explained by the climatic variations of the area, where the cool winter have temperatures below zero. Such condition could be the cause of the reduction in the microbial metabolic activity. On the other hand, during the warmer periods, the microbial population and its activity increase again. The observed seasonal variation is in agreement with several studies that affirm that changes in soil humidity, pH, availability of organic carbon (OC) and input of OM influence the soil biological biomass and therefore its enzymatic activity (Mendham, Sankaran, O'Connell & Grove, 2002; Smithwick, Turner, Metzger & Balser, 2005; Boerner, Brinkman & Smith, 2005;Tabuchi, Kato & Nioh, 2008).
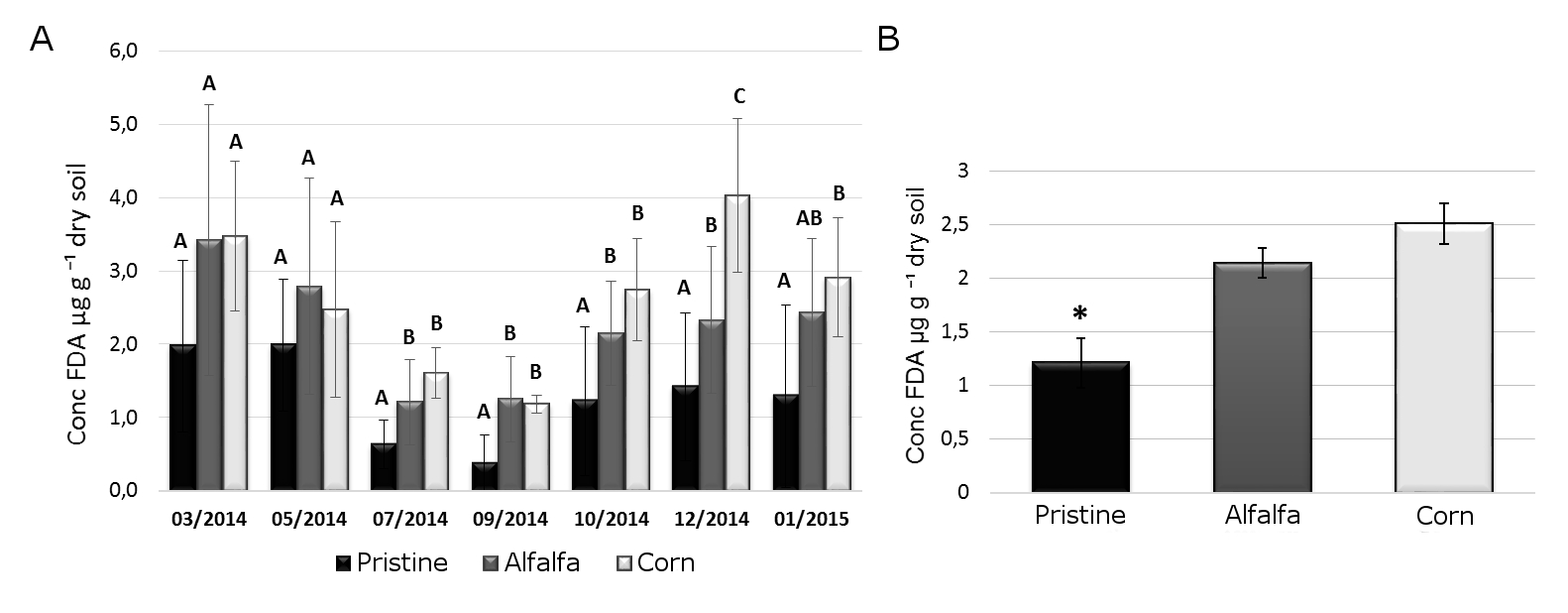
When the three scenarios were compared, significant differences were found among the Pristine soil and the irrigated production systems (Alfalfa and Corn). The Pristine soil (without anthropogenic disturbances nor irrigation) presented significantly lower mean TBA levels (p<0.05) (Figure 1B). This result could be explained by the extreme condition in bare soils. Here, the combination of low humidity, high temperatures and low levels of OM, could negatively influence the TBA. Conversely, Corn and Alfalfa plots presented higher TBA, probably for the positive combination of irrigation and milder soil temperature associated to vegetation cover. In according, Li & Sarah (2015), have reported important increase in the enzymatic activity, including invertase, prolease, catalase, urease and phosphatase produced by the combination of chemical and organic fertilizer, when a “desert becoming to oasis”.
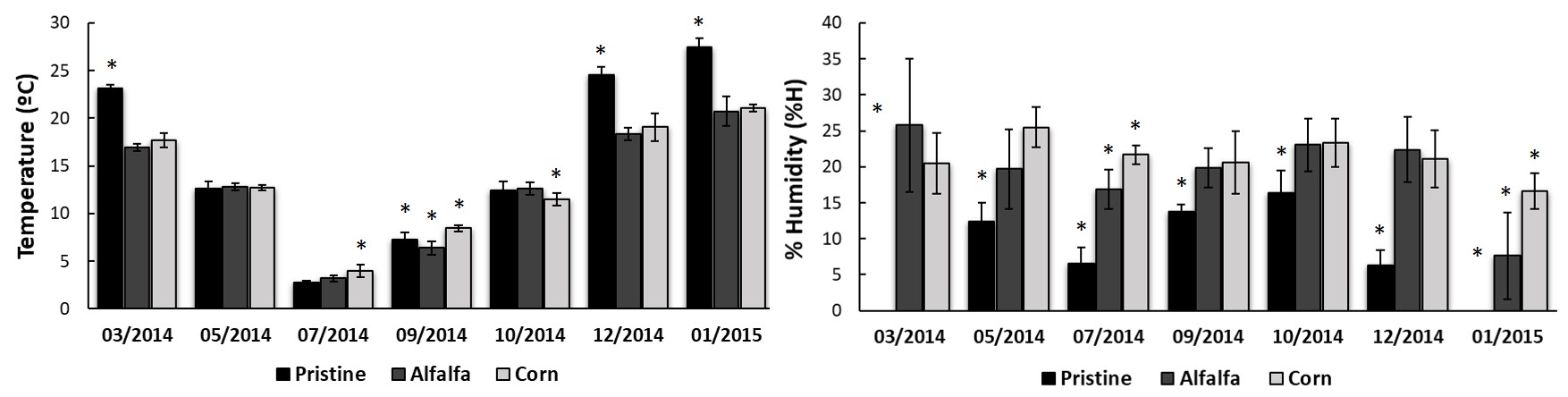
In addition, soil temperature (Figure 2A) and humidity (Figure 2B) were recorded at each point for all samples. The analysis of temperature along the year shows more pronounced variation in the Pristine scene, probably due to lack of protection of the vegetation cover. In managed soils, the variation of temperature along the year was marked, although not as extreme as in the chaparral.
Li & Sarah (2003) and Abbasi, Zafar & Sultan (2010), established that humidity is one of the most influencing environmental variables on the soil biological activity by means of increasing the microbial growth. In this sense, although Pristine soil shared the same overall TBA profile than irrigated soils, it activity was consistently lower all along the year. On the other hand, although pristine soils presented extreme dryness conditions (less than 5% humidity) in January and March (Figure 2B), their microbial activity was measurable and even higher than the observed in the coldest months (Figure 1A). These results would indicate that, in this type of arid ecosystems, the temperature would be more influential than humidity.
Regarding physical-chemical and textural characteristics (Table 1), the Pristine soil showed low contents of clay and TBA levels. In contrast, the soil with corn culture presented higher clay and TBA levels. This is in accordance whit the reported by Burke et al. (1989), pointed that the increase in clay is accompanied by an increase in soil organic carbon (SOC) content, and increased temperatures reduce the SOC levels, probably by microbial decomposition activity. This is especially important in these agroecosystems, since soil mineral composition affects the
formation of biogeochemical interfaces and of a soil-specific microbiota that control the response to introduced organic xenobiotics (Babin et al., 2014).
Considering these results, and the influence of the mineral content on bacterial community, soils with similar textural class were chosen at the moment of selecting the ones to be fingerprinted after DNA extraction (see Table 1).
Soil Microbial Diversity.
Many authors affirm that the shape, size and diversity of microbial community depend on the textural composition (Bach et al., 2010;Furrer Chau, Bagtzoglou & Willig, 2011). Conversely, other authors found that the microbial community size, dynamic and structure, are influenced by irrigation, management practices, soil cover, plant species and application of fertilizers (Drenovski, Vo, Graham, & Scow, 2004; Figuerola et al., 2012; Carbonetto, Rascovan, Alvarez, Mentaberry, & Vázquez, 2014; Agaras, Wall & Valverde, 2014). In this way, permanent vegetation cover, as often occur in grasslands compared with arable soils, results in different community structures and higher concentrations of microbial biomass in the former ecosystem (Zelles, Rackwitz, Yo Bai, Beck & Beese, 1995; Garbeva, van Veen & van Elsas, 2004).
Here, the structure of bacterial community of three different soils was analyzed by DGGE and t-RFLP fingerprint techniques. Two soils under agricultural activity with different crop and irrigation system (Corn and Alfalfa) were compared with a Pristine soil without any management nor irrigation system.
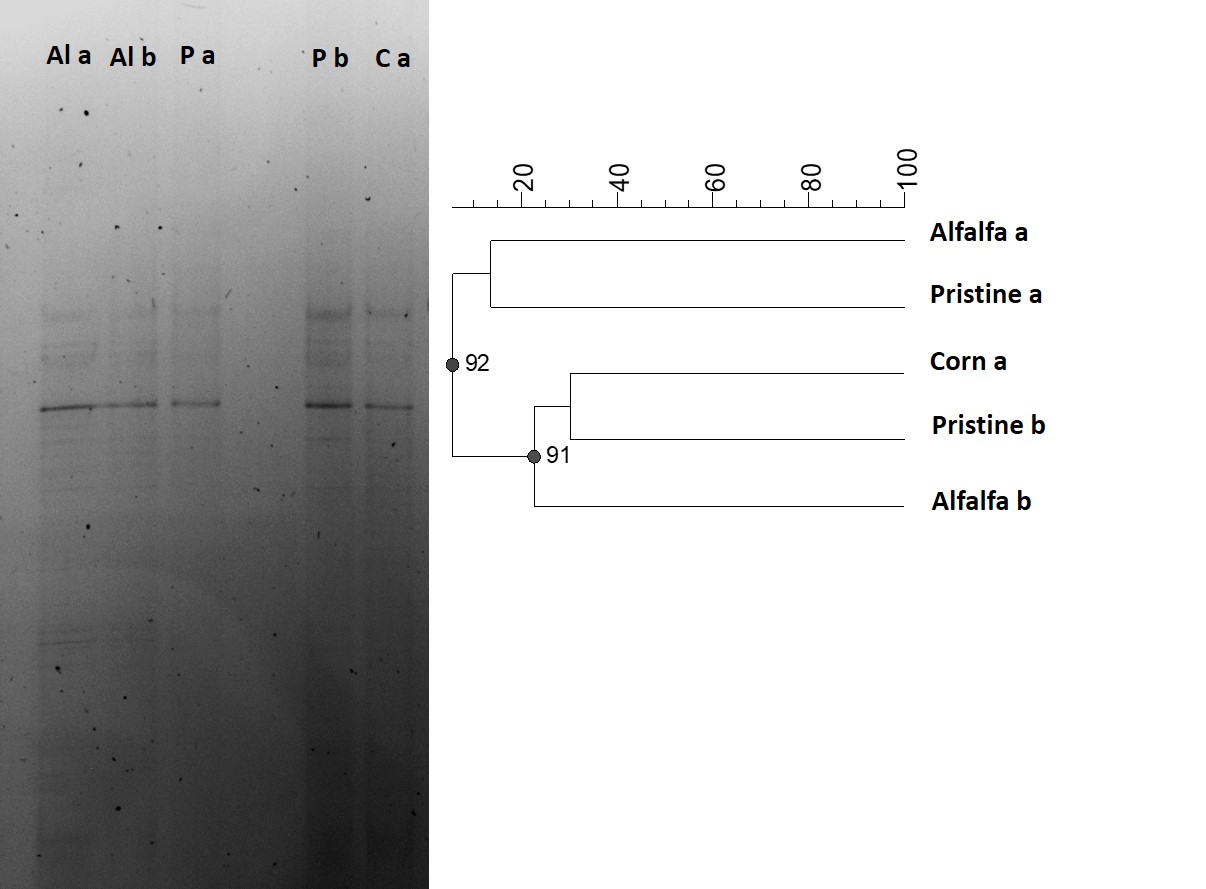
When fingerprint profile of the three environments was analyzed, and against the expected, no differences were found in the structure of the microbial communities. In this way, the Pearson coefficient showed that DGGE banding patterns were very similar among all the soils (Figure 3A) and the corresponding cluster analysis (UPGMA) (Figure 3B) did not reveal- significant differences among plots with different agricultural management nor the Pristine soil. Moreover, Corn soil samples showed more similarities with Pristine soil than the Alfalfa ones, even when both were cultured, fertilized and irrigated plots. Considering the previous reports of many authors showing the marked influence of land use on microbial community structure (Bevivino et al., 2014; Francioli et al., 2014; Naether et al., 2012), the same set of soil samples was reanalyzed and compared by means of T-RFLP.
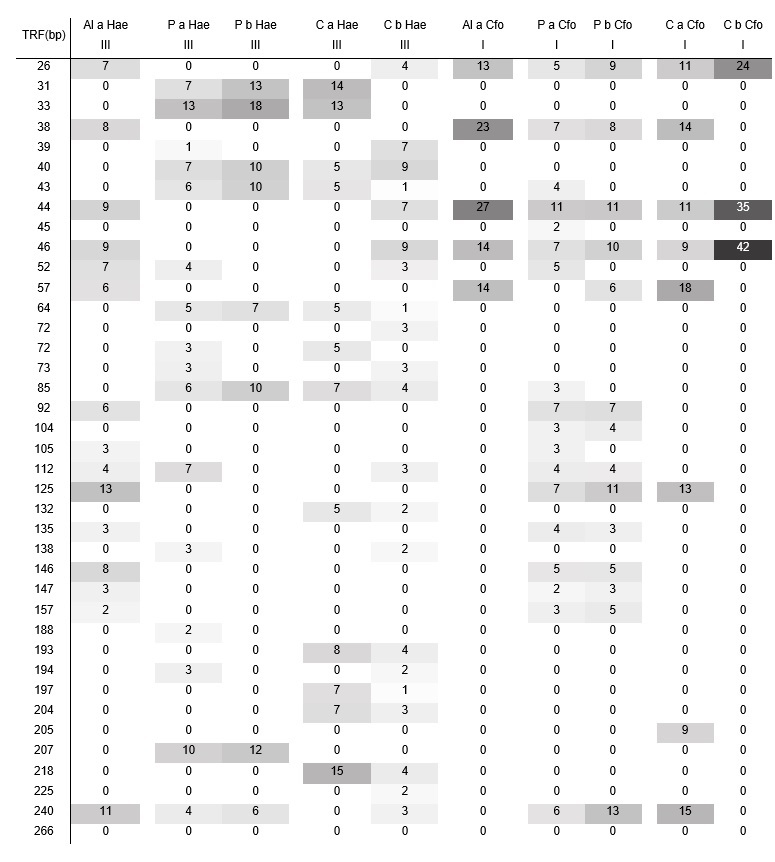
Consistently, the T-RFLP analysis confirmed the lack of differences in microbial biodiversity among the different soils (Table 2), even when the differences in their management practices are evidenced in both, the total biological activity and the physicochemical parameters, especially in the EC and the %TOM.
Although the soil management was not enough to group crops in the same cluster and the biodiversity was low for all soils, its influence was revealed in the richness (R), diversity (H´) and dominance (D) indexes (Table 3). In this way, soil management did not generate significant differences on general bacterial biodiversity but influenced the soil microbial functionality. In general, it was observed that most disturbed and long-term managed plots showed the higher deterioration in soil bacterial community. So, although Corn plots showed the higher biological activity, they had the lowest specific richness (R) and diversity (H´), with high dominance (D) of species. This reflects the impact of management in the bacterial community health, as an ecosystem, reducing soil resilience (Rattan, 1993).
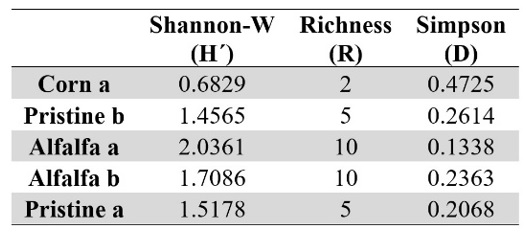
On the other hand, Alfalfa plot showed a good biological activity, and the higher richness and diversity, although the dominance of species was variable.
Even under extreme conditions of water availability and organic matter content (reflected in the low biological activity), the Pristine soils showed higher richness and diversity than Corn plots, with less dominance of species.
Although less frequently, this kind of scenarios were edaphic/climatic characteristics determine the prevalence of dominant bacterial groups and condition the biodiversity over the land use/management, have also been described (Kuramae et al., 2012). However, in our knowledge, these results represent the first evidence, for irrigated agricultural systems, about the prevailing influence of edaphic/climatic conditions over irrigation and land management.
In this context, considering soil microbial activity and diversity are often regarded as an important index of soil ecosystem health and resilience (Garbeva et al., 2004; Torsvik & Ovreas, 2002) these results could be relevant in terms of land management recommendations. In this way, low soil biodiversity could be interpreted as ecosystem vulnerability in terms of topsoil systematization and persistent pesticides input. In addition, the use of inoculants could be an important tool for increasing crop yields.
Conclusions
The results obtained about the soil microbial population structure and its biological activity along the year are significant inputs for moving toward the production maximization with minimal impact on these susceptible environments.
The bacterial biodiversity (in terms of diversity indexes) has remained almost invariable on spite of the long term management of the soils under irrigated agriculture practices, being the soil genesis and environmental condition, principally the temperature, the most influencing factors.
Although farming practices eventually improve the agronomical properties of these arid soils in terms of organic matter content and structure, the bacterial population structure and activity present a hard to change homeostasis. In this sense, despite the use of agrochemical can transform these arid sandy ecosystems into productive irrigated lands, they remain susceptible to pollution associated to the xenobiotics input with low biodegradation potential.
On the other hand, although the techniques used in this work are sensitive and widely employed to detect changes in microbial communities, we consider it important to use next-generation techniques, such as amplicon sequencing of the 16S ribosomal RNA (rRNA) gene method, to identify the global communities of bacteria in these soils.
Finally, based on the obtained preliminary results, an extended assessment of the total microbiota of these soils would be an important input for the decision-making in land management.
Acknowledgments
This work was supported by Instituto Nacional de Tecnología Agropecuaria (PAMSL-1282103: “Gestión de innovaciones para el desarrollo sustentable de la cuenca del Rio Colorado” and PNSUELO 1134043 “Caracterización y funcionalidad de la biota del suelo”).
References
Abbasi, K.M., Zafar, M. & Sultan, T. (2010). Changes in soil properties and microbial indices across various management sites in the mountain environments of Azad Jammu and Kashmir Commun. Soil science and plant analysis. 41, 768-782.
Adam, G. & Duncan, H. (2001). Development of a sensitive and rapid method for the measurement of total microbial activity using fluorescein diacetate (FDA) in a range of soils. Soil biology and biochemistry. 33, 943-951.
Agaras, B.C., Wall, L.G. & Valverde, C. (2014). Influence of agricultural practices and seasons on the abundance and community structure of culturable pseudomonads in soils under no-till management in Argentina. Plant soil. 379: DOI 10.1007/s11104-014-20958.
Bach, E.M., Baer, S.G., Meyer, C.K., Six, J. (2010). Soil texture affects soil microbial and structural recovery during grassland restoration. Soil biology and biochemistry. 42, 2182-2191.
Babin, D., Vogel, C., Zühlke, S., Schloter, M., Pronk, G.J., Heister. K., Spiteller, M., Kögel-Knabner, I., Smalla, K. (2014). Soil mineral composition matters: response of microbial communities to phenanthrene and plant litter addition in long-term matured artificial soils. PLoS ONE. 9, 1-12.
Bevivino, A., Paganin, P., Bacci, G., Florio, A., Pellicer, M.S., Papaleo, M.C., Mengoni, A., Ledda, L., Fani, R., Benedetti, A., Dalmastri, C., 2014. Soil Bacterial Community Response to Differences in Agricultural Management along with Seasonal Changes in a Mediterranean Region. PLoS ONE. 9(8): e105515. doi:10.1371/journal.pone.0105515
Boerner, R.E.J., Brinkman, J.A. & Smith, A. (2005). Seasonal variations in enzyme activity and organic carbon in soil of a burned and unburned hardwood forest. Soil Biology & Biochemistry. 37, 1419–1426.
Bouyoucos, G., 1962 Hydrometer method improved for making particle size analysis of soils. Agronomy Journal. 54, 464-465.
Burke, I.C., Yonker, C.M., Parton, W.J., Cole, C.V., Flach, K., Schimel, D.S. (1989). Texture, Climate, and Cultivation Effects on Soil Organic Matter Content in U.S. Grassland Soils. Soil Science Society of America Journal. 53, 800-805.
Carbonetto, B., Rascovan, N., Alvarez, R., Mentaberry, A., Vázquez, M.P. (2014). Structure, composition and metagenomic profile of soil microbiomes associated to agricultural land use and tillage systems in Argentine pampas. PLoS ONE. 9(6): e99949. doi:10.1371/journal.pone.0099949.
del Valle, H.F. (1998). Patagonian soils: a regional synthesis. Ecología Austral. 8, 103-123.
Drenovski, R.E., Vo, D., Graham, K.J., Scow, K.M. (2004). Soil water content and organic carbon availability are major determinants of soil microbial community composition. Microbial Ecology. 48, 424–430.
Entry, J.A., Mills, DE., Mathee, K., Jayachandran, K., Sojka, RE., Narasimhan, G. (2008). Influence of irrigated agriculture on soil microbial diversity. Applied Soil Ecology. 40, 146-154.
Enwall, K. & Hallin, S. (2009). Comparison of T-RFLP and DGGE techniques to assess denitrifier community composition in soil. Letters in Applied. Microbiology. 48, 145–148.
Evans, R.G. & Sadler, E.J. (2009). Methods and technologies to improve efficiency of water use. Water Resources Research. 44. doi:10.1029/2007WR006200.
Fang, M., Kremer, R.J., Motavalli, P.P., Davis, G. (2005). Bacterial diversity in rhizospheres of nontransgenic and transgenic corn. Applied and Environmental Microbiology. 71, 4132–4136.
Figuerola, E.L.M., Guerrero, L.D., Rosa, S.M., Simonetti, L., Duval, M.E., Galantini, J.A., Bedano, J.C., Wall, L.G., Erijman, L. (2012). Bacterial indicator of agricultural management for soil under no-till crop production. PLoS ONE. 7(11): e51075. doi:10.1371/journal.pone.0051075.
Francioli, D., Ascher, J., Ceccherini, M.T., Pietramellara, G. (2014). Land use and seasonal effects on a Mediterranean soil bacterial community. Journal of Soil Science and Plant Nutrition. 14, 710-722.
Furrer Chau J., Bagtzoglou A.C. & Willig M.R. (2011). The effect of soil texture on richness and diversity of bacterial communities. Environmental. Forensics, 12, 333–341.
Gabriel, J.L., Almendros, P., Hontoria, C., Quemada, M., 2012. The role of cover crops in irrigated systems: Soil salinity and salt leaching. Agriculture, Ecosystems & Environment. 158, 200-207.
Garbeva, P., van Veen, J.A. & van Elsas, J.D. (2004). Microbial Diversity in Soil: Selection of Microbial Populations by Plant and Soil Type and Implications for Disease Suppressiveness. Annual Review of Phytopathology. 42, 243–70.
Green, V.S., Stott, E.D. & Diack, M. (2006). Assay for fluorescein diacetate hydrolytic activity: Optimization for soil samples. Soil Biology & Biochemistry. 38, 693–701.
Griffiths, B.S., Bonkowski, M., Roy, J. & Ritz, K. (2001). Functional stability, substrate utilization and biological indicators of soils following environmental impacts. Applied Soil Ecology. 16, 49–61.
Hamarashid, N.H., Othman, M.A. & Hussain, M.A.H. (2010). Effects of soil texture on chemical compositions, microbial populations and carbon mineralization in soil. Egyptian journal of experimental biology. 6, 59 – 64.
Kuramae, E.E., Yergeau, E., Wong, L.C., Pijl, A.S., van Veen, J.A., Kowalchuk, G.A. (2012). Soil characteristics more strongly influence soil bacterial communities than land-use type. FEMS Microbiology Ecology. 79, 12–24.
Li, X. & Sarah, P., 2003. Enzyme activities along a climatic transect in the Judean Desert. Catena. 53, 349-363.
Matson, P.A., Parton, W.J., Power, A.G., Swift, M.J. (1997). Agricultural Intensification and Ecosystem Properties. Science. 277, 504-509.
Meena, V.S., Maurya, B.R., Meena, R.S., Meena, S.K., Singh, N.P., Malik, V.K., Kumar, V., Jat, L.K. (2014). Microbial dynamics as influenced by concentrate manure and inorganic fertilizer in alluvium soil of Varanasi, India. African Journal of Microbiology Research. 8, 257-263.
Mendham, D.S., Sankaran, K.V., O'Connell, A.M., Grove, T.S. (2002). Eucalyptus globulus harvest residue management effects on soil carbon and microbial biomass at 1 and 5 years after plantation establishment. Soil Biology and Biochemistry. 34, 1903-1912.
Mohd-Aizat, A., Mohamad-Roslan, M.K., Sulaiman W.N.A., Karam, D.S. (2014). The relationship between soil pH and selected soil properties in 48 years logged-over forest. International Journal of Environmental Science. 4, 1129-1140.
Naether, A., Foesel, B.U., Naegele, V., Wüst, P.K., Weinert, J., Bonkowski, M., Alt, F., Oelmann, Y., Polle, A., Lohaus, G., Gockel, S., Hemp, A., Kalko, E.K.V., Linsenmair, K.E., Pfeiffer, S., Renner, S., Schöning, I., Weisser, W.W., Wells, K., Fischer, M., Overmann, J., Friedricha, M. W. (2012). Environmental Factors Affect Acidobacterial Communities below the Subgroup Level in Grassland and Forest Soils. Applied and Environmental Microbiology. 78, 7398–7406.
Panigatti, J.L. (2010). Argentina: 200 años, 200 suelos. Ediciones INTA, Buenos Aires.
Pérez, A.J., Abrahao, R., Causape, J; Cirpka, O.A., Burger, C.M. (2011). Simulating the transition of a semi-arid rainfed catchment towards irrigation agriculture. Journal of Hydrology. 409, 663-681.
Rattan L. (1993). Tillage effects on soil degradation, soil resilience, soil quality, and sustainability. Soil and Tillage Research. 27, 1-8.
Scanlon, B.R., Jolly, I., Sophocleous, M., Zhang, L. (2007). Global impacts of conversions from natural to agricultural ecosystems on water resources: Quantity versus quality. Water Resources Research. 43, 1-18.
Schnürer, J. & Rosswall, T. (1982). Fluorescein diacetate hydrolysis as a measure of total microbial activity in soil and litter. Applied and Environmental Microbiology. 43, 1256-1261.
Singh, B.K., Munro, S., Reid, E., Ord, B., Potts, J.M., Paterson, E., Millard, P. (2006). Investigating microbial community structure in soils by physiological, biochemical and molecular fingerprinting methods. European Journal of Soil Science. 57, 72–82.
Smith, C.J., Danilowicz, B.S., Clear, A.K., Costello, F.J., Wilson, B., Meijer, W.G. (2005). T-Align, a web-based tool for comparison of multipleterminal restriction fragment length polymorphism profiles. FEMS Microbiology Ecology. 54, 375–380.
Smithwick, E.H.A., Turner, M.G., Metzger, K.L., Balser, T.C. (2005). Variation in NH4+ mineralization and microbial communities with stand age in lodgepole pine (Pinus contorta) forests, Yellowstone National Park (USA). Soil Biology and Biochemistry. 37, 1546-1559.
Solaiman, Z. & Marschner, P. (2007) DGGE and RISA protocols for microbial community analysis in soil. In: Advanced techniques in soil microbiology (Varma, A. and Oelmüller, R., eds.), Springer-Verlag, Berlin Heidelberg, 167–180.
Şumalan, R., Alexa, E., Negrea, M., Doncean, A. (2010). Comparative study on biological activity and edaphic microflora composition for four soil types from SDE Timisoara. Research Journal of Agricultural Sciences. 42, 324-327.
Tabuchi, H., Kato, K. & Nioh, I. (2008). Season and soil management affect soil microbial communities estimated using phospholipid fatty acid analysis in a continuous cabbage (Brassica oleracea var. capitata) cropping system. Soil Science and Plant Nutrition. 54, 369-378.
Tilman, D. (1999). Global environmental impacts of agricultural expansion: The need for sustainable and efficient practices. Proceedings of the National Academy of Sciences. USA. 96, 5995–6000.
Torsvik, V. & Ovreas, L. (2002). Microbial diversity and function in soil: from genes to ecosystems. Current Opinion in Microbiology. 5, 240–245
Walkley, A. & Black, I.A. (1934). An examination of the Degtjareff method for determining soil organic matter, and proposed modification of the chromic acid titration method. Soil Science. 37, 29-38.
Weisburg, G.W., Barns, S.M., Pelletier, D.A. Lane, D.J. (1991). 16S ribosomal DNA amplification for phylogenetic study. Journal of Bacteriology. 173, 697-703.
You, Y., Wang, J., Huang, X., Tang, Z., Liu, S., Sun, O.J. (2014). Relating microbial community structure to functioning in forest soil organic carbon transformation and turnover. Ecology and Evolution. 4, 633-647.
Zelles, R., Rackwitz, O., Yo Bai, T., Beck, I., Beese, F. (1995). Discrimination of microbial diversity by fatty acid profiles of phospholipids and lipopolysaccharides in differently cultivated soils. Plant Soil. 170, 115-122.
Zhao, J., Ni, T., Li, Y., Xiong, W., Ran, W., Shen, B., Shen, Q., Zhang, R. (2014). Responses of bacterial communities in arable soils in a rice-wheat cropping system to different fertilizer regimes and sampling times. PLoS ONE. 9(1): e85301. doi:10.1371/journal.pone.0085301.
Ziesemer, K., Mann, A., Sankaranarayanan, K. et al. (2015). Intrinsic challenges in ancient microbiome reconstruction using 16S rRNA gene amplification. Scientific Reports. 5, 16498. https://doi.org/10.1038/srep16498
Duncan, D. B. (1955). Multiple Range and Multiple F Tests. Biometrics. 11(1), 1–42. https://doi.org/10.2307/3001478

